Method Article
Modified Most Probable Number Assay to Quantify Salmonella in Raw and Ready-to-Cook Chicken Products
In This Article
Summary
Accurately quantifying Salmonella in poultry at low levels is a current industrial and regulatory challenge. This protocol describes an MPN assay that enables quantification of Salmonella in raw and ready-to-cook poultry products. This method is fast, sensitive, and aligns with FSIS guidelines, enhancing food safety and supporting public health efforts.
Abstract
Salmonella is a leading cause of foodborne illness in the United States, particularly in poultry products. Traditional methods for detecting Salmonella focus on prevalence rather than quantification, which limits their utility in assessing contamination levels and risks. This study introduces a novel most probable number (MPN) assay designed to quantify Salmonella in ready-to-cook poultry products, such as chicken cordon bleu. The method involves washing the poultry sample, concentrating the rinse through centrifugation, and serially diluting it in a 48-well block. The MPN assay is integrated with the loop-mediated isothermal amplification (LAMP) method to provide a sensitive, accurate, and rapid quantification of Salmonella contamination within the same timeframe as existing Food Safety and Inspection Service (FSIS) protocols. Results show a strong linear correlation between the MPN-LAMP measurements and theoretical inoculation levels (R² = 0.933). However, variability at lower concentrations highlights challenges in accurately detecting Salmonella at these levels, with the practical lower limit of detection estimated at approximately 300 CFU/g. Potential refinements to improve the protocol's applicability include increasing the quantity sampled to further improve the limit of detection, optimizing enrichment media formulations, and expanding molecular detection to target multiple Salmonella serovars. Overall, this study presents a practical tool for the food industry, enabling reliable quantification of Salmonella contamination in poultry products, contributing to improved food safety and public health.
Introduction
As a leading cause of foodborne illness, hospitalization, and death in the U.S., Salmonella has a significant public health and economic impact. The pathogen's estimated economic burden in 2013 alone was $3.67 billion1. Although recent regulatory initiatives aim to reduce salmonellosis by 25% by 20302, gaps in current detection and mitigation strategies remain evident, particularly in aligning processing plant surveillance with public health outcomes3 .
Frozen ready-to-cook poultry products, which have been implicated in multiple Salmonella outbreaks, are a significant concern for public health. In response, the Food Safety and Inspection Service (FSIS) classified Salmonella as an adulterant in these products. Currently, FSIS Microbiology Laboratory Guidebook (MLG) 4.15 focuses solely on determining the prevalence of Salmonella in poultry products4. Under this guideline, collected samples are enriched for 18-24 h and then screened using the Molecular Detection System (MDS), which identifies the presence or absence of Salmonella but does not offer insight into the level of contamination. While this approach is valuable for detecting the presence of pathogens, it fails to provide quantitative information that could help food processors assess contamination risks more accurately and take targeted corrective actions.
In this study, we developed a method to augment detection from prevalence to quantification of microbial pathogens. It was designed for seamless integration into existing processes to detect Salmonella in poultry products with minimal disruption to current FSIS protocols. Instead of simply enriching the bulk sample, the method begins by washing the poultry products using media consistent with current FSIS methods. The rinse is then distributed into the first column of a 48-deep well block. Serial dilutions are performed across the remaining five columns, and the block is incubated for 18-24 h, aligning with the MLG 4.15 protocol. After incubation, the wells are tested for Salmonella, and the results are used to calculate the most probable number (MPN)5,6. This approach allows for quantification of contamination within the same time frame as the current FSIS process, making it a practical option for both industry and regulatory use. Figure 1 depicts a block diagram summarizing the modified MPN assay. The figure includes photographs taken at specific steps, the 48-well block utilized for dilution and growth of replicates, and the three techniques used as benchmarks to assess the most probable number of Salmonella present in ground chicken. In the first phase of this study, we utilized irradiated ground chicken to minimize the impact of background microflora and uncertainty of measurements relative to verified inoculum before applying the protocol to non-irradiated chicken samples.
Protocol
NOTE: All work associated with this protocol should be conducted within a Biosafety Level 2 (BSL-2) laboratory. When appropriate, this protocol should be conducted within a biological safety cabinet (BSC) to maintain aseptic conditions and minimize the risk of sample contamination or operator exposure to microbial pathogens. When transferring samples outside the BSC, use sealed containers to maintain sample integrity and prevent spillage in case of accidental drops. Preferably, disposable components should be used throughout the procedure to mitigate the possibility of cross-contamination. In cases where disposables are not feasible, ensure all equipment and materials are sterile prior to use. Proper waste management is crucial; all used disposable components should be discarded as biohazard waste. Autoclave reusable materials before reuse to ensure proper sterilization and containment of potentially hazardous materials. Adhering to these precautions not only safeguards sample integrity but also minimizes the risk of operator exposure to microbial pathogens.
1. Preparation of meat samples
- Acquiring and processing meat samples
- Fresh meat
- Acquire ground chicken from the fresh meat department of local retailers. Transfer all samples to storage at 4 °C and process within 24 h after receipt. Aseptically divide meat into 25 g samples.
- Vacuum seal and irradiate the sample. Here, Texas A&M AgriLife National Center for Electron Beam Research irradiated meat subjected to a dose of ~25 kGy.
NOTE: While irradiation was used as a control measure in this study to ensure the elimination of background microflora, it is not a prerequisite for the protocol in practical applications as shown by the non-irradiated ready-to-cook products used in the subsequent section. In field settings, alternative methods such as selective media or specificity of molecular diagnostics can address potential interference from non-target microorganisms.
- Ready-to-cook chicken products
- Acquire ready-to-cook chicken products from the frozen food section of local retailers. Aseptically divide into 25 g samples.
- Collect samples from the center of the individual pieces to ensure all ingredients (e.g., breading and cheese) are included.
- Fresh meat
- Media preparation
- Prepare Buffered Peptone Water (BPW) by dissolving 25 g of BPW powder in 1 L of nanopure H2O.
- Prepare brain heart infusion (BHI) plates. To do this, dissolve 37 g of BHI powder in 1 L of nanopure H2O and add 15 g of agar into BHI solution. Sterilize all media by autoclaving at 121 °C for 15 min. Pour 20 to 25 mL of media into Petri Dishes with Clear Lid (100 mm x 15mm).
NOTE: Its best to pour the plates within a biological safety cabinet to maintain aseptic conditions.
2. Cell culture
- Prepare the initial culture by streaking Salmonella enterica serovar Typhimurium ATCC 14028 on a BHI agar plate and incubate at 37 °C overnight.
- Prepare overnight cultures by inoculating 25 mL of BHI broth with one colony of freshly grown Salmonella. Aerobically grow cultures overnight at 37 °C with shaking at 100 rpm.
3. Inoculation of poultry samples
- Culture dilution and plating
- Prepare a series of 10-fold dilutions of the overnight culture in BPW to achieve final concentrations of approximately 1 x 108 to 1 x 101 CFU/mL. Assume the concentration of the Salmonella overnight culture is 1 x 109 CFU/mL.
- Transfer 0.5 mL of the overnight culture into 4.5 mL of BPW, mix, then transfer 0.5 mL of the dilution into 4.5 mL of BPW for each additional dilution.
- Spread 10 µL of the 1 x 103 CFU/mL dilution on a BHI agar plate in triplicate for cell enumeration to calculate the overnight culture concentration.
- Inoculation of meat samples
- Aseptically transfer 25 g of irradiated ground chicken into a sterile stomacher bag in duplicate. The bag is 7.5 x 12 inches and is labeled as 1.63 L. The bag contains a filter partition with a hole diameter of 330 µm, and there are 285 per square cm.
- Inoculate each sample with 1 mL of the target concentration dilution. For example, add 1 mL of culture from the 1 x 10³ CFU/mL dilution to achieve a level of contamination of approximately 1,000 cells/25 g chicken. Gently distribute the liquid inoculum over the surface of the chicken samples using a sterile cell spreader and allow it to stand for 1 h at 4 °C.
- Prepare negative control samples by adding 1 mL of sterile BPW.
4. Sample processing
- Add 225 mL of BPW to each sample. The ratio of volume to media was selected to align with FSIS MLG 4.154.
- Homogenize the samples using the Stomacher7 at normal speed and a 120 s duration.
- Centrifugation and resuspension
- Carefully remove the liquid from the filtered side of the bag using a 50 mL pipette. Split the liquid into two sterile centrifuge bottles.
- Balance the bottles with sterile BPW to ensure equal weights. Centrifuge the samples at 10,000 x g for 10 min. Discard the supernatant. Resuspend the cell pellet in 3 mL of BPW broth with a sterile spatula.
- Add an additional 27 mL of BPW broth and mix thoroughly by stirring with a spatula. Combine the contents of both centrifuge bottles into one bottle for each sample.
5. MPN block set-up
NOTE: Table 1 depicts a schematic of the dilutions in a 48-well block.
- Add 3 mL of the resuspended sample to each well in column 1 of the 48-well block (8 replicates).
- Prepare a series of 10-fold dilutions across columns 1-6 within the block using an eight-channel pipette.
- Add 0.3 mL of sample into 2.7 mL of BPW pipette to mix. Repeat for each dilution. Incubate blocks overnight (~18 h) at 37 °C with shaking at ~100 rpm.
6. Plating and enumeration
- Modified drop plate enumeration
- Plate 7 µL of overnight grown sample of each dilution in a 4 x 6 grid on an agar plate using a multichannel pipette (Figure 2). Using a 4 x 6 grid on two plates better accommodates 8 samples, as opposed to the typical 6 x 6 grid of droplets8.
- Allow plates to air dry for 10 min before incubation. Incubate the agar plates overnight (~18-24 h) at 37 °C. After incubation, count the number of colonies on each plate.
7. qPCR detection of Salmonella
- DNA extraction using a commercial kit
- Mix cultures in the 48-well block by pipetting up and down several times. Pipette 200 µL of each culture into a 96-well PCR plate.
- Seal and then centrifuge the plate at 6,600 x g for 10 min. Remove the supernatant and add 20 µL of the kit reagent to the pellet.
- Resuspend the pellet by pipetting up and down. Seal and heat the plate at 99 °C for 10 min, followed by cooling to 20 °C.
- Centrifuge again at 6,600 x g for 10 min. Use 2 µL of the supernatant for qPCR analysis.
- Plate setup
- Prepare the qPCR reaction mixture according to established protocol9 as follows: 10 µL of 2x Master Mix; 0.4 µL of each primer and probe (10 µM working solution): invA forward: 5'-GTTGAGGATGTTATTCGCAAAGG-3', invA reverse: 5'-GGAGGCTTCCGGGTCAAG-3', invA probe: 5'-CCGTCAGACCTCTGGCAGTACCTTCCTC-3' labeled with the Cal Fluor Orange 560 fluorenes dye; 0.2 µL of internal amplification control (IAC) template (6 x 104 copy/µL)9; 0.4 µL of each IAC primer and probe (10 µM): IAC forward: 5'-GGCGCGCCTAACACATCT-3', IAC reverse: 5'-TGGAAGCAATGCCAAATGTGTA-3', IAC probe: 5'-TTACAACGGGAGAAGACAATGCCACCA-3' labeled with TAMRA dye. Adjust volume with ddH2O to 20 µL total.
- Perform real-time PCR with the following cycling conditions9: 95 °C for 10 min (initial denaturation of DNA and activation of hot-start polymerase), 40 cycles of 95 °C for 15 s and 60 °C for 1 min, use default Ct settings to export results for analysis.
8. Detection using 3M MDS assay
- Follow the molecular detection assay Salmonella kit protocol. Mix cultures in the 48-well block by pipetting up and down several times. Pipette 20 µL of each sample into the kit-provided lysis tube.
- Heat the samples at 100 °C for 15 min. The solution will turn from pink to yellow. Incubate the samples for 10 min at room temperature. The solution will change from yellow to pink.
- Transfer 20 µL of lysate into a reagent tube and load the reagent tubes into the holder.
- Add the holder to the MDS instrument and configure the software to communicate the information about the kit and the sample. The instrument requires each well to be labeled with the lot number for the assay and a sample name. Run the MDS software and export the report.
9. Data analysis
- Classification of positive and negative results.
- For 4 x 6 drop plating, assess spots on agar plates with at least 1 colony as positive and spots on agar plates with no growth as negative.
- For qPCR, assess wells that have a Ct less than or equal to 30 as positive and wells that have a Ct greater than 30 as negative.
- For MDS, use the results from the MDS system, reported as positive or negative.
- MPN calculation
- Analyze annotated positives and negatives using the simple maximum probability resolution (SMPR) method previously described6 or alternative verified MPN calculators.10
Results
Irradiated meat
In regression analysis, a slope of 1 indicates that for every unit increase in the independent variable (x-axis), the dependent variable (y-axis) increases by exactly 1 unit. This suggests a proportional relationship between the two variables, meaning that the change in the dependent variable mirrors the change in the independent variable. An intercept of 0 means that when the independent variable is 0, the dependent variable is also 0. This suggests that there is no fixed offset or bias in the relationship between the two variables. Together, a slope of 1 and an intercept of 0 suggest agreement between the variables11,12. This ideal scenario indicates no systematic error or bias and would be the expected outcome in a well-calibrated system where measurements align perfectly with predictions.
Figure 3A presents the MPN-LAMP assay (y-axis) plotted against the theoretical inoculum verified by the spread plate analysis discussed in the note in step 3.1.1. (x-axis). The regression line equation, y = -72.36 + 0.161x, with an R² value of 0.978, indicates a strong linear correlation between the MPN-LAMP results and the inoculated bacterial levels. The high R² value suggests that the MPN-LAMP assay is highly reliable in the quantitative prediction of bacterial contamination. However, the slope of 0.161 deviates significantly from the ideal slope of 1, indicating that the MPN-LAMP method underestimates the inoculum, predicting only 16.1% of bacterial concentration in the theoretical inoculum. Additionally, the negative intercept of -72.36 suggests a negative bias at lower concentrations, further deviating from the ideal.
Figure 3B illustrates the MPN-qPCR method (y-axis) plotted against the theoretical inoculum (x-axis). The regression line, y = -46.84 + 0.0395x, with an R² value of 0.978, demonstrates a strong linear relationship between the MPN-qPCR assay and the theoretical inoculum. Although the high R² indicates a robust correlation, the slope of 0.0395 is even smaller than that of the MPN-LAMP results, suggesting a greater degree of underestimation in bacterial contamination by MPN-qPCR. The negative intercept of -46.84 further indicates a potential downward bias at lower concentrations.
Figure 3C presents the MPN-plating results (y-axis) plotted against the theoretical inoculum (x-axis), with a regression equation of y = -330.4 + 0.3138x and an R² value of 0.944. While this R² value indicates a strong linear relationship, it is slightly lower than those of the MPN-LAMP and MPN-qPCR results, suggesting greater variability in colony quantification. The slope of 0.3138, though higher than the slopes observed in the other methods, still indicates an underestimation of bacterial concentration relative to the theoretical inoculum. Additionally, the negative intercept of -330.4 points to considerable bias, especially at lower concentrations, which may indicate potential inaccuracies in plating techniques or sample handling. This slight reduction in R² and the variability observed could indicate that the traditional plating process is more variable than the molecular methods.
Comparison methods
All three methods (MPN-LAMP, MPN-plating, and MPN-qPCR) showed a tendency to underestimate bacterial levels relative to the theoretical inoculation. This underestimation can be attributed to cell recovery issues, where Salmonella may not fully detach from the chicken surface or particulate matter during processing, reducing its transfer into the rinse or assay. Since the Salmonella was directly inoculated onto the chicken, a portion of bacterial cells may bind strongly to the surface, limiting recovery in the rinse step. Additionally, some cells may remain attached to particulate matter in the pellet after centrifugation, further lowering recovery. Approaches such as enzymatic treatments, mechanical processing, or optimized stomaching protocols could enhance bacterial detachment. These strategies are discussed further in the discussion section, emphasizing their potential for improving recovery rates
Figure 4A presents the recovery on the y-axis against three MPN methods on the x-axis. The percent recovery, calculated as the ratio of the MPN value to the theoretical inoculation level, yielded the following results: the MPN-LAMP method had a mean recovery of 15.19% with a standard deviation of 10.04%; the MPN-plating method had a mean recovery of 13.13% with a standard deviation of 11.45%; and the MPN-qPCR method had a mean recovery of 6.67% with a standard deviation of 3.19%. A Tukey-Kramer HSD test did not identify a statistically significant difference in recovery between the methods (P > 0.1969), suggesting that all three methods performed similarly in terms of recovery when processing Salmonella-inoculated chicken.
Despite identical treatment across methods for separating Salmonella from the chicken matrix, variations in detection among MPN-LAMP, MPN-qPCR, and MPN-plating were observed. This suggests that factors beyond the physical separation of bacteria from the sample may impact detection. The presence of inhibitors in the sample matrix could interfere with the molecular methods-qPCR and LAMP-affecting their sensitivity and overall performance. Additionally, bacterial viability may have impacted the ability of Salmonella to grow on solid agar plates (MPN plating) versus liquid broth (qPCR, LAMP), leading to variation in recovery between the methods.
To further explore the sensitivity of the methods, we first examined the number of positive responses at each dilution for MPN-LAMP, MPN-qPCR, and MPN-plating across seven independent trials (Table 2). At lower dilutions (1 x 10-5 and 1 x 10-4), MPN-LAMP demonstrated a higher number of positive responses compared to both MPN-plating and MPN-qPCR, suggesting that the LAMP method is more sensitive to low concentrations of Salmonella. When samples were more concentrated (dilutions less than 1 x 10-3), all three methods showed an increase in positive detections, with convergence occurring around the 1 x 10-2 dilution, where all methods detected a similar number of positive samples.
Figure 4B presents the percentage of positive measurements at each dilution across the three methods, providing a direct comparison of their sensitivity in detecting Salmonella. The percentage of positive responses offers a clearer visualization of method performance across the dilution series, and error bars representing ± 95% Clopper-Pearson confidence intervals (CI) provide insight into variability at each level. At lower dilutions, the wider confidence intervals indicate greater variability in the detection performance of each method. Consistent with the raw number of positive detections, MPN-LAMP detects a higher percentage of positive samples at the most dilute concentrations (1 x 10-5), while MPN-qPCR and MPN-plating detect fewer positives. As the bacterial concentration increases, the methods converge, with all three approaches showing close to 100% detection at higher concentrations (e.g., 1 x 10-2 and above). The narrowing of the confidence intervals at these higher concentrations indicates greater consistency across all methods, emphasizing their comparable performance when bacterial levels are higher.
The findings in Figure 3 suggest that the consistent underprediction (quantitatively) of all three methods could be driven by incomplete separation of Salmonella from the sample matrix. However, molecular inhibitors or bacterial viability factors could be driving the observed variation in detection at different dilutions (qualitatively) and across different methods (qualitatively and quantitatively). The tighter confidence intervals at higher concentrations in Figure 4B indicate less variability and more consistent qualitative performance across methods, while the broader intervals at lower concentrations point to increased uncertainty in qualitative detection.
Detection in ready-to-cook chicken
We selected the LAMP method for the detection of Salmonella in ready-to-cook chicken as it displayed a greater number of positive samples (14.3%) than other methods tested at the lowest Salmonella dilution. At the lowest Salmonella dilution (1 x 10-5), none of the samples were detected as positive by the in-house qPCR method. On the other hand, some positives were detected by plating (7.1%, half the number detected by LAMP). However, the results from this culture-based method may be affected when moving to more complex products such as ready-to-cook chicken, where the presence of background microflora could skew the results because samples are not irradiated. For these reasons, we decided to investigate the MPN-LAMP method for quantification of Salmonella from ready-to-cook chicken.
Figure 5 depicts the results of the MPN-LAMP assay used to quantify Salmonella in inoculated, frozen, ready-to-cook chicken cordon bleu samples. The theoretical level of inoculation (CFU/g) was plotted on the x-axis, while the corresponding MPN-LAMP measurements were plotted on the y-axis. The regression analysis yielded the equation y = -15.04 + 0.3869x, with a coefficient of determination (R²) of 0.933, indicating a strong linear correlation between the theoretical inoculum and the MPN-LAMP results. This high R² value suggests that the MPN-LAMP method reliably quantifies Salmonella in these samples, although the slope of 0.3869 indicates that the method underestimates the actual inoculum level. The shaded area around the regression line represents the 95% confidence interval.
To better illustrate the performance of MPN-LAMP at lower inoculation levels, the inset graph in Figure 5 zooms in on data points below 300 CFU/g. Based on the regression equation, the lower limit of detection can be estimated at approximately 40 CFU/g (38.9 CFU/g). However, the 95% confidence interval suggests that the practical lower limit of detection is closer to 325 CFU/g, as any negative MPN-LAMP values should be interpreted as zero. The red line in the inset represents zero, and although no data points fell below this line, the potential for values to drop below zero, particularly at lower inoculum levels, introduces the possibility of false-negative results. This trend highlights the difficulties of accurately quantifying Salmonella at low concentrations, and additional data would be necessary to definitively establish the lower limit of detection.

Figure 1: Block Diagram of the Modified MPN Assay for Salmonella Detection. A schematic overview of the modified Most Probable Number (MPN) assay. It outlines key steps in the process, starting with the inoculation of irradiated ground chicken samples, followed by sample homogenization, centrifugation, aliquot distribution, and incubation under specified conditions. The figure includes photographs of the 48-well block utilized for dilution and replicate growth, along with three techniques used to produce positive and negative results for the Simple Maximum Probability Resolution (SMPR) calculation. Please click here to view a larger version of this figure.

Figure 2: Schematic of the drop plate assay. The drop plate assay setup for a specific dilution (e.g., undiluted, 10-1, 10-2, etc.) from the 48-well block. Each sample is plated in a 4 x 6 grid format on an agar plate using a multichannel pipette. The layout was modified from the traditional 6 x 6 grid to two 4 x 6 grids to ensure six replicates from eight dilutions could be accommodated. Please click here to view a larger version of this figure.
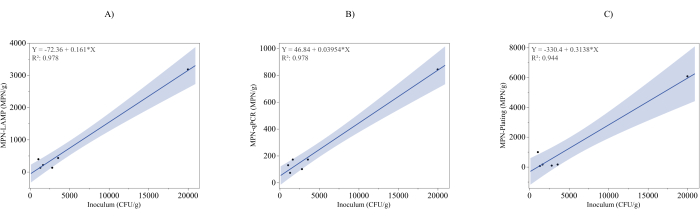
Figure 3: Comparison of Inoculum with MPN methods. (A) A plot of MPN-LAMP concentration (y-axis) against the theoretical inoculum verified by plating (x-axis). A fitted regression line (y = -72.36 + 0.161x) with a coefficient of determination (R² = 0.978) demonstrates a strong linear relationship. (B) A plot of MPN-qPCR concentration (y-axis) against the theoretical inoculum (x-axis), with a fitted regression line (y = -46.84 + 0.0395x) and R² = 0.978, showing a similarly strong correlation. (C) A plot of MPN-plating concentration (y-axis) against the theoretical inoculum (x-axis), with a regression line (y = -330.4 + 0.3138x) and R² = 0.944, indicating a strong but slightly lower correlation than LAMP and qPCR. The regression equation and the coefficient of determination for each figure are presented as an insert in the top left corner of the graphs. Please click here to view a larger version of this figure.

Figure 4: Comparison of MPN methods. (A) The figure presents the percent recovery, calculated as the ratio of the MPN value to the theoretical inoculation level for MPN-LAMP, MPN-plating, and MPN-qPCR methods. The MPN-LAMP method had a mean recovery of 15.19% (±10.04%), the MPN-plating method had a mean recovery of 13.13% (± 11.45%), and the MPN-qPCR method had a mean recovery of 6.67% (± 3.19%). The box plots highlight the distribution of the recovery data, with outliers indicated. Error bars represent the range of data within 1.5 times the interquartile range (IQR). (B) Percentage of positive Salmonella detections at each dilution across three methods: MPN-LAMP (blue), MPN-plating (red), and MPN-qPCR (green). The x-axis represents the dilution levels (10-6 to 10), and the y-axis shows the percentage of positive measurements. Error bars indicate the ± 95% Clopper-Pearson confidence intervals, which reflect the variability in detection for each method. MPN-LAMP shows higher percentages of detections than other methods at lower dilutions, while all methods converge near 100% detection at higher concentrations. Please click here to view a larger version of this figure.
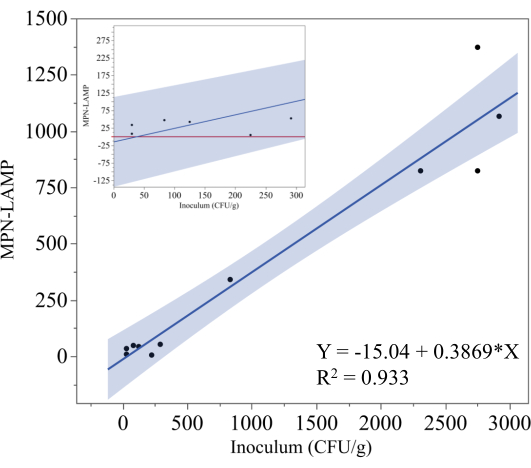
Figure 5: MPN method for ready-to-cook chicken. Regression analysis of MPN-LAMP results against the theoretical inoculation level of Salmonella in frozen, ready-to-cook chicken cordon bleu samples. The regression line is represented by y = -15.04 + 0.3869x with an R² of 0.933, indicating a strong linear relationship. The shaded region represents the 95% confidence interval. The inset focuses on the MPN-LAMP measurements at lower inoculation levels. Please click here to view a larger version of this figure.
| 1 | 2 | 3 | 4 | 5 | 6 | |
| A | n=1 | n=1 | n=1 | n=1 | n=1 | n=1 |
| B | 2 | 2 | 2 | 2 | 2 | 2 |
| C | 3 | 3 | 3 | 3 | 3 | 3 |
| D | 4 | 4 | 4 | 4 | 4 | 4 |
| E | 5 | 5 | 5 | 5 | 5 | 5 |
| F | 6 | 6 | 6 | 6 | 6 | 6 |
| G | 7 | 7 | 7 | 7 | 7 | 7 |
| H | 8 | 8 | 8 | 8 | 8 | 8 |
| 100 | 10-1 | 10-2 | 10-3 | 10-4 | 10-5 |
Table 1: Schematic of Replicates and Dilutions in a 48-Well Block. This figure illustrates the arrangement of replicates and serial 10-fold dilutions within the 48-well block. Each column represents a specific dilution factor, ranging from, 100 to 10-5 with technical replicates for each dilution distributed across the eight rows.
| Dilution | ||||||
| 10-5 | 10-4 | 10-3 | 10-2 | 10-1 | 100 | |
| MPN-LAMP | 8 | 16 | 31 | 3 | 56 | 56 |
| MPN-plating | 4 | 11 | 32 | 55 | 56 | 56 |
| MPN-qPCR | 0 | 7 | 26 | 55 | 56 | 56 |
Table 2: Number of positive events. The table displays the number of positive reactions (over 56 total replicates). The results are arranged into a table where columns represent dilution, and rows are representative of the method.
Discussion
Significance of the protocol
Salmonella remains a major concern in food safety, particularly within poultry products, which are often implicated in foodborne illness outbreaks13,14. As a leading cause of bacterial foodborne illness in the United States, reliable methods for detecting Salmonella in both fresh and ready-to-cook poultry products are critical to ensuring food safety15. The ability to quantify Salmonella at low levels is vital for identifying contamination that may otherwise go undetected. Method sensitivity is particularly important for regulatory compliance and public health, as even low doses of Salmonella can result in illness, especially in vulnerable populations.
The MPN-LAMP assay utilizes the MDS system, which is currently used by FSIS in MLG 4.15, to determine the prevalence of contamination4. The assay presented here provides an extension of the current FSIS method to quantify viable Salmonella, even at low levels, providing more precise data on the level of contamination. Results from this study demonstrated a strong correlation between the MPN-LAMP method and theoretical inoculation values, with an R² of 0.933 (Figure 5). This suggests the method is reliable for quantifying Salmonella in food, although some underestimation occurred at lower inoculation levels.
Critical steps
The protocol for Salmonella quantification necessitates precision in sample preparation, inoculation, and molecular analysis. Proper aseptic technique, accurate dilutions, and adherence to incubation conditions are pivotal for reliable detection and enumeration of Salmonella. Ensuring sterile conditions during the transfer of poultry samples and media is essential to prevent contamination.
Equally important is the homogenization of the samples during stomaching, followed by careful resuspension of the pellet following centrifugation to separate the bacteria from the food matrix. Proper handling during these steps ensures that viable cells are recovered for further analysis. Accurate pipetting during the serial dilution process for MPN setup, coupled with consistent incubation conditions, is crucial for obtaining reproducible results.
Finally, molecular detection methods, such as qPCR, require precision and reproducibility in DNA extraction and qPCR cycling to avoid false negatives. Although DNA extraction is inherently variable, standardized reagents and procedures ensure repeatability by minimizing inhibitor interference and improving DNA recovery. These steps are fundamental for confirming the presence of Salmonella and for targeting specific genetic markers that can help overcome limitations in traditional plating and enrichment methods.
Potential pitfalls and limitations
Bacterial viability
One of the primary challenges of implementing a growth-based assay such as the MPN-plating is bacterial viability, and this is especially relevant for complex ready-to-cook chicken samples. Frozen, breaded, and stuffed chicken products, such as chicken cordon bleu, are particularly prone to temperature fluctuations and extended storage periods, both of which can stress Salmonella cells and make the pathogen harder to detect in growth-based assays. In addition, the presence of non-chicken ingredients (e.g., cheese, breading, ham, spices, butter) introduces further complexity in the food matrix and can inhibit the growth of Salmonella during enrichment phases, complicating accurate quantification. Additionally, Salmonella must compete against background microbes in culture-based methods16, which is especially relevant for ready-to-cook samples with multiple ingredients which adds diversity to the microbiota relative to raw chicken. Finally, handling and processing stressors, such as antimicrobials, can influence Salmonella recovery17.
Bacterial viability may have impacted the performance of the methods tested in this study. All methods underestimated Salmonella concentrations at low levels and had higher detection variability at lower pathogen concentrations (Figure 4A,B). However, the molecular-based MPN-LAMP method outperformed MPN-plating and MPN-qPCR in qualitative detection at lower inoculation levels. The results emphasize the need for refinement at low pathogen concentrations.
Sample heterogeneity
Another important limitation involves the heterogeneity of food samples, which can introduce variability in Salmonella detection results. The protocol was designed to disperse 30 mL of homogenized sample, allowing for the distribution of 8 replicates as 3 mL aliquots (24 mL total, 6 mL excess) across the block. While this conservative approach (resulting in an extra sample) accounts for potential losses, it introduces subsampling as the entire volume is not tested.
The heterogeneity of the food sample can affect measurement accuracy when transferring samples from the redispersed mixture into the block and performing dilutions18,19. According to the Theory of Sampling (TOS), fundamental sampling errors can be reduced by increasing the sample mass or reducing compositional heterogeneity through particle size reduction20. However, while some studies indicate that breaking down the matrix by stomaching does not significantly impact bacterial recovery from surface contamination, reducing the particle size of the food matrix has been shown to decrease bacterial recovery in some cases21,22,23. In this study, the particle size was reduced by stomaching prior to enrichment; while this may reduce sampling error by reducing compositional heterogeneity in samples, stomaching and centrifugation may impact bacterial recovery.
Finally, the heterogeneous composition and morphology of the food, as well as whether the contamination is internal or on the surface, may lead to variability in pathogen recovery. Previous work18,22 has demonstrated that strong bacterial binding to tissue surfaces can reduce recovery rates. Studies where enzymes such as trypsin, collagenase, and endopeptidase were used to release bacteria from meat and poultry matrices24,25 and pectinase and cellulase to leafy greens19 demonstrate the potential to improve Salmonella recovery from food matrices.
Suggestions for troubleshooting
To prevent issues, ensure that all media and reagents are freshly prepared and sterile. Expired or improperly sterilized materials may lead to contamination or affect the growth of Salmonella. If contamination is observed, verify that the aseptic technique was consistently followed throughout the protocol, particularly during sample inoculation, dilution, aliquoting, and plating.
When colony counts are lower than expected, verifying the accuracy of serial dilutions and inoculum concentrations can help identify errors. Incubation conditions, including temperature, time, and agitation, must be carefully monitored to support optimal Salmonella growth. In cases of low or no growth, inadequate stomaching or leakage from bags could contribute to uneven distribution of the bacteria.
If molecular detection produces inconsistent results, checking DNA extraction procedures and verifying the quality of primers and probes are key steps. Also, variability between food matrices may present different levels of inhibition, which can be monitored using the internal amplification control (IAC). Proper storage and handling of extraction reagents are crucial to prevent false negatives due to degradation or contamination. More efficient DNA extraction methods may also be warranted.
Incorporating treatments such as the GentleMACS system26, which employs mechanical, thermal, and enzymatic actions, can further optimize bacteria recovery from food. This system can reduce the heterogeneity of the food matrix, making it easier to transfer uniform samples to the block and carry out accurate serial dilutions.
Potential refinement and expansion
There are several opportunities for refining and expanding this protocol. Exploring alternative enrichment media formulations could improve the recovery of stressed or sublethally injured Salmonella cells, particularly from complex food matrices such as chicken cordon bleu. While selective agar is commonly used to target Salmonella, cells that are injured or stressed due to processing and storage conditions may lead to false negatives on such media. To overcome this challenge, optimizing the balance between selective pressure and microbial recovery could enhance the detection of Salmonella serovars that are less competitive under standard enrichment conditions. This refinement could improve overall assay sensitivity, particularly in challenging food products where microbial stress is common. Shortening incubation times could accelerate the time to results, though this may reduce sensitivity and limit detection. Future work should explore advancements in enrichment media to support rapid bacterial replication, ensuring accurate quantification while meeting stakeholder requirements.
Further, investigating different incubation temperatures and durations for the MPN assay could optimize Salmonella recovery from various poultry products. This could expand the protocol's applicability to other foodborne pathogens that require distinct growth conditions, thereby broadening its utility in food safety testing. Automation could enhance the scalability of this protocol by integrating robotic liquid handlers for aliquoting and dilution steps. Continuous-flow centrifuges or high-capacity centrifuge systems may also address batch processing limitations, enabling high-throughput applications in industrial settings.
Expanding the molecular detection aspect to include multiplex PCR could enable simultaneous detection of multiple pathogens, improving throughput in surveillance and testing environments. Additionally, incorporating next-generation sequencing (NGS) could provide valuable serotype and virulence information directly from MPN enrichment cultures, facilitating epidemiological investigations and source-tracking efforts.
Lastly, the limit of quantification/detection can be expanded by increasing the quantity of samples used in the MPN. For example, if the weight of the meat sample was increased from 25 g to 325 g, then the limit of detection could be extrapolated to 4 CFU/g using the linear regression equation. However, assuming the same relative 95% confidence interval, the practical lower limit of quantification is >30 CFU/g.
Conclusion
The development of the MPN-LAMP assay in this study offers a valuable and practical tool for quantifying Salmonella contamination in poultry products. With a strong correlation to theoretical inoculation levels (R² = 0.933), the method reliably detects Salmonella at low levels, which is critical for food safety and regulatory compliance. This capability is especially important in detecting contamination that may otherwise go unnoticed, allowing producers to proactively address potential risks. The MPN-LAMP assay provides a quantitative alternative to traditional prevalence-based methods, offering a level of Salmonella contamination in a variety of poultry products, including more complex ready-to-cook items.
One of the key strengths of this method lies in its adaptability to different food matrices, particularly frozen and processed products like chicken cordon bleu, where traditional methods might struggle due to the presence of additional ingredients such as breading and cheese. The integration of molecular techniques, such as LAMP and qPCR, with the MPN assay ensures a comprehensive approach to detection, overcoming challenges posed by background microbes and processing conditions. These molecular methods enhance the sensitivity of the protocol, making it a versatile tool in routine food safety testing and contamination surveillance, with the potential to reduce the risk of Salmonella-related outbreaks.
While no single detection method is without limitations, the MPN-LAMP assay demonstrates consistent performance across a range of inoculum levels and food matrices. Variability at lower inoculation levels can be addressed through future refinements, such as optimizing sample preparation and media formulations to maximize recovery, but this does not detract from the overall robustness of the protocol. By providing accurate detection of Salmonella cells, even under challenging conditions, the MPN-LAMP method equips the food industry with an invaluable tool for safeguarding public health.
Looking ahead, there is room for further enhancement, such as expanding the assay to include more Salmonella serovars or incorporating next-generation sequencing for greater pathogen characterization. Nonetheless, this protocol already represents a significant advancement in Salmonella detection and quantification, providing a practical, efficient, and reliable method that meets the growing demands of food safety testing.
Disclosures
All the authors declare that there is no conflict of interest.
Acknowledgements
This research was supported by the U.S. Department of Agriculture, Agricultural Research Service (USDA-ARS), National Program 108, Current Research Information System numbers 8072-42000-093-000-D and 8072-42000-094-000-D. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U. S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Materials
| Name | Company | Catalog Number | Comments |
| 48 deep well block 4.6ml | Fisher Scientific International, Inc | NC1964628 | |
| Agar - Solidifying Agent (Difco) | Becton, Dickinson and Company (BD) | 281230 | |
| Analytical Balance | Mettler Toledo | JL602-G/L | Equipment |
| Analytical Balance | Mettler Toledo | AB54-S | Equipment |
| Autoclave - Amsco Lab250, Laboratory Steam Sterilizer | Steris plc | LV-250 | Equipment |
| Biological Safety Cabinet, Type A2, Purifier Logic+ | Labconco Corporation | 302411101 | Equipment |
| Brain Heart Infusion (BHI) Broth | Becton, Dickinson and Company (BD) | 237500 | |
| Buffered Peptone Water | Bio-Rad Laboratories Inc. | 3564684 | |
| Cell Spreader - L-shaped | VWR | 76208-438 | |
| Centrifuge Microcentrifuge 5424 | Eppendorf | 5424 | Equipment |
| Centrifuge, Avanti J-25 | Beckman Coulter, Inc. | Equipment | |
| DNA Extraction - PreMan Ultra Sample Preparation Reagent | Thermo Fisher Scientific Inc. | 4318930 | |
| Ground Chicken | Local retailers | ||
| IAC forward primer: 5'-GGCGCGCCTAACACATCT-3' | Integrated DNA Technologies | ||
| IAC probe: 5'-TTACAACGGGAGAAGACAATGC CACCA-3' labeled with 5' TAMRA/3' BHQ-2 | Biosearch Technologies | ||
| IAC reverse primer: 5'-TGGAAGCAATGCCAAATGTGTA-3' | Integrated DNA Technologies | ||
| Incubator - Inova 4230 incubator shaker | New Brunswick Scientific | 4230 | Equipment |
| Inoculating Loop - Combi Loop 10µL and 1µL | Fisher Scientific International, Inc | 22-363-602 | |
| invA forward primer: 5'-GTTGAGGATGTTATTCGCAAAG G-3' | Integrated DNA Technologies | ||
| invA probe: 5'-CCGTCAGACCTCTGGCAGTAC CTTCCTC-3' labeled with 5' Cal Fluor Orange 560/3' BHQ-1 | Biosearch Technologies | ||
| invA reverse primer: 5'-GGAGGCTTCCGGGTCAAG-3' | Integrated DNA Technologies | ||
| Irradiation Treatment | Texas A&M Agrilife Research National Center for Electron Beam Research | Service | |
| Luria Bertani (LB) Broth | Becton, Dickinson and Company (BD) | 244620 | |
| Manual pipette Pipet-Lite LTS Pipette L-1000XLS+ | Mettler Toledo | 17014382 | Equipment |
| Manual pipette Pipet-Lite LTS Pipette L-100XLS+ | Mettler Toledo | 17014384 | Equipment |
| Manual pipette Pipet-Lite LTS Pipette L-10XLS+ | Mettler Toledo | 17014388 | Equipment |
| Manual pipette Pipet-Lite LTS Pipette L-200XLS+ | Mettler Toledo | 17014391 | Equipment |
| Manual pipette Pipet-Lite LTS Pipette L-20XLS+ | Mettler Toledo | 17014392 | Equipment |
| Manual pipette Pipet-Lite Multi Pipette L8-200XLS+ | Mettler Toledo | 17013805 | Equipment |
| Manual pipette Pipet-Lite Multi Pipette L8-20XLS+ | Mettler Toledo | 17013803 | Equipment |
| Media Storage Bottle -PYREX 1L Square Glass Bottle, with GL45 Screw Cap | Corning Inc. | 1396-1L | Equipment |
| Media Storage Bottle -PYREX 2L Round Wide Mouth Bottle, with GLS80 Screw Cap | Corning Inc. | 1397-2L | Equipment |
| Microtiter plate, 96 well plate, flat bottom, polystyrene, 0.34cm2, sterile, 108/cs | MilliporeSigma | Z707902 | |
| Mixer - Vortex Genie 2 | Scientific Industries Inc. | SI-0236 | Equipment |
| Molecular Detection Assay 2-Salmonella kit | Neogen | MDA2SAL96 | |
| Molecular Detection Instrument | Neogen | MDS100 | Equipment |
| Motorized pipette controller, PIPETBOY2 | INTEGRA Biosciences Corp. | 155019 | Equipment |
| PCR Mastermix 2× TaqMan Gene Expression | Thermo Fisher Scientific Inc. | 4369542 | |
| Petri Dish Rotator - bioWORLD Inoculation Turntable | Fisher Scientific International, Inc | 3489E20 | Equipment |
| Petri Dishes with Clear Lid (100 mm x 15mm) | Fisher Scientific International, Inc | FB0875713 | |
| Pipette Tips GP LTS 1000µL S 768A/8 | Mettler Toledo | 30389273 | |
| Pipette Tips GP LTS 20µL 960A/10 | Mettler Toledo | 30389270 | |
| Pipette Tips GP LTS 200µL F 960A/10 | Mettler Toledo | 30389276 | |
| Ready to cook chicken products | Local retailers | ||
| Reagent Reservoir, 25 mL sterile reservoir used with multichannel pipettors | Thermo Fisher Scientific Inc. | 8093-11 | |
| Realtime PCR - 7500 Real-Time PCR system | (Applied Biosystems, Foster City, CA) | 2750036476 | Equipment |
| Serological Pipettes, Nunc Serological Pipettes (10 mL) | Thermo Fisher Scientific Inc. | 170356N | |
| Serological Pipettes, Nunc Serological Pipettes (2 mL) | Thermo Fisher Scientific Inc. | 170372N | |
| Serological Pipettes, Nunc Serological Pipettes (25 mL) | Thermo Fisher Scientific Inc. | 170357N | |
| Serological Pipettes, Nunc Serological Pipettes (50 mL) | Thermo Fisher Scientific Inc. | 170376N | |
| Spreader - Fisherbrand L-Shaped Cell Spreaders | Fisher Scientific International, Inc | 14-665-230 | |
| Stomacher bag, Nasco Whirl-Pak Write-On Homogenizer Blender Filter Bags | Thermo Fisher Scientific Inc. | 01-812 | |
| Stomacher 80 Biomaster Lab Blender | Seward | 30010019 | Equipment |
| Thermocycler (GeneAmp PCR system 9700) | Applied Biosystems | 535028293 | Equipment |
| Water Filtration - Elga Veolia Purelab Flex | Elga LabWater | PF2XXXXM1-US | Equipment |
| Whirlpak bags 1.63L | VWR | 11216-777 |
References
- Batz, M., Hoffmann, S., Morris, J. G. Disease-outcome trees, eq-5d scores, and estimated annual losses of quality-adjusted life years (qalys) for 14 foodborne pathogens in the united states. Foodborne Pathogens and Disease. 11 (5), 395-402 (2014).
- . The Grand Challenge: Salmonella Available from: https://tellus.ars.usda.gov/stories/articles/the-grand-challenge-salmonella (2024)
- National Advisory Committee on Microbiological Criteria in Foods (NACMCF). Response to questions posed by the food safety and inspection service: Enhancing Salmonella control in poultry products. J Food Prot. 82 (4), 645-668 (2019).
- Food Safety and Inspection Service. . 4.15 Isolation and identification of Salmonella from meat, poultry, pasteurized egg, siluriformes (Fish) products and carcass and environmental sponges. , (2024).
- Irwin, P., Reed, S., Brewster, J., Nguyen, L., He, Y. P. Non-stochastic sampling error in quantal analyses for campylobacter species on poultry products. Analytical and Bioanalytical Chemistry. 405 (7), 2353-2369 (2013).
- Irwin, P., Tu, S., Damert, W., Phillips, J. A modified gauss-newton algorithm and ninety-six well micro-technique for calculating mpn using excel spreadsheets. Journal of Rapid Methods & Automation in Microbiology. 8 (3), 171-191 (2000).
- Ravishankar, S., Ahmed, E. Y., Carlstrom, C. Food microbiology: A laboratory manual. Food Microbiology. 21, 489 (2004).
- Chen, C. Y., Nace, G. W., Irwin, P. L. A 6 x 6 drop plate method for simultaneous colony counting and mpn enumeration of campylobacter jejuni, listeria monocytogenes, and escherichia coli. J Microbiol Methods. 55 (2), 475-479 (2003).
- Suo, B., He, Y., Tu, S. I., Shi, X. A multiplex real-time polymerase chain reaction for simultaneous detection of salmonella spp., escherichia coli o157, and listeria monocytogenes in meat products. Foodborne Pathogens and Disease. 7 (6), 619-628 (2010).
- Jarvis, B., Wilrich, C., Wilrich, P. T. Reconsideration of the derivation of most probable numbers, their standard deviations, confidence bounds and rarity values. J Appl Microbiol. 109 (5), 1660-1667 (2010).
- Stevens, R., Poppe, K. Validation of clinical prediction models: What does the "calibration slope" really measure. Journal of Clinical Epidemiology. 118, (2019).
- Miller, M. E., Hui, S. L., Tierney, W. M. Validation techniques for logistic regression models. Statistics in Medicine. 10 (8), 1213-1226 (1991).
- Galán-Relaño, &. #. 1. 9. 3. ;., et al. Salmonella and salmonellosis: An update on public health implications and control strategies. Animals. 13 (23), 3666 (2023).
- Gorski, L., et al. Growth assessment of salmonella enterica multi-serovar populations in poultry rinsates with commonly used enrichment and plating media. Food Microbiology. 119, 104431 (2024).
- Schmidt, J. W., et al. Evaluation of methods for identifying poultry wing rinses with salmonella concentrations greater than or equal to 10 cfu/ml. J Food Prot. 87 (11), 100362 (2024).
- Gorski, A., Liang, L. S. Effect of enrichment medium on real-time detection of salmonella enterica from lettuce and tomato enrichment cultures. Journal of Food Protection. 73 (6), 1047-1056 (2010).
- Guillén, S., Nadal, L., Álvarez, I., Mañas, P., Cebrián, G. Impact of the resistance responses to stress conditions encountered in food and food processing environments on the virulence and growth fitness of non-typhoidal salmonellae. Foods. 10 (3), 617 (2021).
- Rohde, A., Hammerl, J. A., Appel, B., Dieckmann, R., Al Dahouk, S. Sampling and homogenization strategies significantly influence the detection of foodborne pathogens in meat. BioMed Research International. 2015, (2015).
- Wang, D., Wang, Z., He, F., Kinchla, A. J., Nugen, S. R. Enzymatic digestion for improved bacteria separation from leafy green vegetables. Journal of Food Protection. 79 (8), 1378-1386 (2016).
- Pitard, F. F. . Theory of sampling and sampling practice. , (2019).
- Sharpe, A. . in Detecting pathogens in food. , 52-68 (2003).
- Hannah, J., et al. Effect of stomaching on numbers of bacteria recovered from chicken skin. Poultry Science. 90 (2), 491-493 (2011).
- Mcmeekin, T., Thomas, C. Retention of bacteria on chicken skin after immersion in bacterial suspensions. Journal of Applied Bacteriology. 45 (3), 383-387 (1978).
- Rodrigues-Szulc, U., Ventoura, G., Mackey, B., Payne, M. Rapid physicochemical detachment, separation and concentration of bacteria from beef surfaces. Journal of Applied Bacteriology. 80 (6), 673-681 (1996).
- Vibbert, H. B., et al. Accelerating sample preparation through enzyme-assisted microfiltration of salmonella in chicken extract. Biotechnol Prog. 31 (6), 1551-1562 (2015).
- Armstrong, C. M., et al. Use of a commercial tissue dissociation system to detect salmonella-contaminated poultry products. Analytical and Bioanalytical Chemistry. 416 (3), 621-626 (2024).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved