Rapid Amplification of cDNA Ends
Overview
Source: Pablo Sanchez Bosch2, Sean Corcoran2 and Katja Brückner1,2,3
1Eli and Edythe Broad Center of Regeneration Medicine and Stem Cell Research
2Department of Cell and Tissue Biology,
3Cardiovascular Research Institute, University of California San Francisco, San Francisco, CA, USA
Rapid Amplification of cDNA Ends (RACE) is a technique that allows amplification of full-length cDNA from mRNA by extending to the 3’ or 5’ end, even without prior knowledge of the sequence (Frohman et al., 1988). In contrast to regular PCR, it uses only one specific PCR primer and a second non-specific primer that will indiscriminately bind to most mRNAs. Depending on whether the area to be amplified is on the 3’ or 5’ end of the mRNA, the second primer is chosen to bind the polyA tail (3’ end) or a synthetic linker added to all transcripts (5’ end). These primer combinations are known to yield “one-sided” or “anchored” PCR due to PCR amplification using the known sequence of one side- 3’ or 5’(Ohara et al., 1989). The approach allows the capture of gene-specific rare mRNAs that otherwise would be hard to detect, e.g. because of their relatively low expression or unknown complete sequence (Frohman et al., 1988).
RACE is initiated with a reverse transcription step (RT) to synthetize single stranded complimentary DNA (cDNA) from mRNA. This is followed by two consecutive PCRs that amplify the cDNA fragments from a gene of interest. To perform RACE, the sequence of the gene of interest must be at least partially known, as it is needed to design the gene-specific primers (GSPs; Frohman, 1994; Liu and Gorovsky, 1993). As the second primer pair is a generic primer that will anneal to all transcripts present in the sample, the specificity of the PCR reactions is reduced. RACE is therefore ideally performed with two nested PCRs to lower the chances of amplifying non-specific transcripts (Figure 1). Depending on the direction of amplification relative to the GSP, the technique is categorized as 5’ or 3’ RACE (Figure 1).
3’ RACE (Figure 1A) takes advantage of the poly(A) tail present in mRNAs as a generic binding site for the non-specific primer. This simplifies the technique, as one can synthetize cDNA by using oligo (dT) primers. GSPs are located in the 5’ region of the transcripts. This RACE variant allows the detection of transcript variants and different 3’ UTRs (Scotto Lavino et al., 2007a). In 5’ RACE (Figure 1B), GSPs are located in the 3’ region of the gene. To be able to use a non-specific primer that binds to all transcripts, an adapter that serves as generic primer binding site is attached to the 5’ RNA ends. To attach the adapter to the mRNA, the 5’ cap that protects mRNAs against exonucleases and promotes translation must be removed (Bird et al., 2016). Opposed to 3’ RACE, 5’ RACE helps finding differential 5’ splice variants and alternative 5’ UTRs (Scotto Lavino et al., 2007b).
Here we will perform 3’ RACE to identify and isolate different transcript variants using the known 5’ sequence (Figure 2A) encoded by the Drosophila dSmad2 (Smox) gene. dSmad2, a fly Smad protein, is an important transducer of Activin-β signaling, a pathway of the TGF-β superfamily. dSmad2 regulates multiple cellular processes, such as cell proliferation, apoptosis and differentiation (Upadhyay et al., 2017). As differentially spliced transcripts of dSmad2 may have different functions in the adult fly, a first step in exploring these potential functions is to assess all transcript variants of dSmad2 at this developmental stage.
Procedure
1. Experiment set up for 3’ RACE
- Design a 5’ specific primer for a gene of interest (gene specific primer 1, GSP-1). It must be highly specific, as it is the only primer that introduces specificity. Therefore, a relatively long primer would be preferable (typical length around 24 nucleotides, Tm ranging from 55-65˚C).
- Design a second, nested primer (GSP-2), i.e. the primer should be located 3’ of GSP-1. This step is optional, but performing a second PCR with a nested primer will increase the yield and specificity of the PCR for the gene of interest. The primers used to amplify dSmad2 cDNAs are listed in Table 2.
2. Synthesis of cDNA
- Extract RNA from samples, using an RNA extraction method or kit following the manufacturer’s protocol. Treat the samples with DNase to avoid amplification of genomic DNA. In our example, we extracted RNA from 10 whole adult flies.
- Measure the RNA concentration using a microvolume spectrophotometer and adjust with RNase free water to a maximum of 100 ng/μl if needed.
- Prepare cDNA in a reverse transcription (RT) reaction, using a kit recommended for RACE. Prepare a master mix, with the following reagents per sample:
- 4 μl Reverse transcription buffer (5x)
- 2 μl Oligo(dT)20 primer (an oligo consisting of 20 deoxythymidines)
- 10 μl RNA (max. 1 μg)
- 3 μl ddH2O
- 1 μl reverse transcriptase
- TOTAL: 20 μl - Incubate the reaction for 90 min at 42ºC. Include a negative control omitting reverse transcriptase.
- Inactivate the reverse transcriptase by incubating at 85ºC for 5 min.
- In order to dilute DNA levels for efficient PCR, add 80 μl TE buffer to bring the total volume to 100 μl.
3. Amplification of cDNA fragments
- Prepare the first-round amplification PCR mix by using a high-fidelity (HF) Taq polymerase:
- 4 μl 5x HF Taq polymerase buffer
- 0.4 μl dNTP mix (10 mM each)
- 1 μl GSP1 primer (10 μM)
- 1 μl Oligo(dT)20 primer (10 μM)
- 1 μl cDNA template
- 0.2 μl HF Taq DNA polymerase
- 12.4 μl ddH2O
- Total: 20 μl - Run the PCR by using the PCR1 program (Table 1). Include a negative control using as template the original RNA product without RT. Further, a ‘no primer’ control may be included.
- Optional: nested PCR to increase specificity and the amount of cDNA products. Dilute 1 μl of the PCR products and negative control 1:20 in TE buffer. Use them as template for the second PCR reactions:
- 4 μl 5x HF Taq polymerase buffer
- 0.4 μl dNTP mix (10 mM each)
- 1 μl GSP2 primer (10 μM)
- 1 μl Oligo(dT)20 primer (10 μM)
- 1 μl PCR product template
- 0.2 μl HF Taq DNA polymerase
- 12.4 μl ddH2O
- Total: 20 μl - Run the second PCR using the PCR2 program (Table 1)
4. Isolation of cDNA fragments
- Prepare a 1-2% agarose gel with TAE buffer, using ethidium bromide or an alternative DNA stain.
- Mix 5 μl of sample with 1 μl of 6x gel loading buffer.
- Load the samples and 1kb DNA ladder as marker and run the gel at 120 V for about 45 min or until the dye front is ~75% of the way down the gel. Ensure the samples do not run out of the gel.
- Check the gel bands under a UV lamp. For further processing of the products, use low intensity UV, locate the amplified bands and cut them from the gel using a scalpel.
- Purify the DNA from the gel by using a method like freeze-squeeze or a kit.
- Store the purified cDNA fragments at -20ºC or use them immediately for further applications, such as sequencing or cloning.
Results
There are two annotated transcripts for Drosophila dSmad2 in FlyBase (Fig. 2A). Our results, however, reveal three different transcripts for dSmad2, ranging in size from 750 bp to 1400 bp (Fig. 2B). Differences in the intensity of the bands indicate their relative expression levels. Among the predicted products, one transcript is predominant (Fig. 2B, black arrow A), and one is expressed at a lower level (Fig. 2B, black arrow B). In addition, a previously undescribed smaller product was detected (Fig. 2B, grey arrow C).
The identification of such rare transcripts is only possible with a sensitive method such as RACE. Following RACE, one can clone the fragments obtained by PCR to study the transcript sequence, find its similarity with other transcript variants and clone it for transgenesis. Such experiments help investigate the functions of transcript variants specific to a tissue or developmental stage.
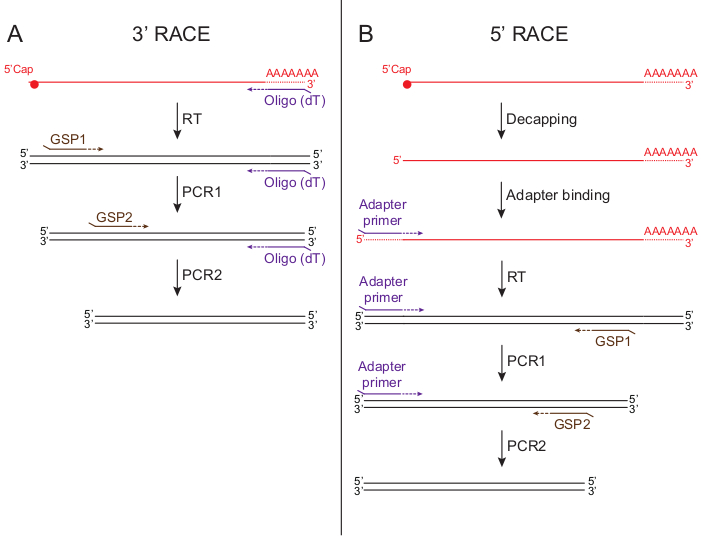
Figure 1. Schematics of the two different RACE approaches. A) 3’ RACE. cDNA is synthesized by using oligo (dT) primers. The same oligo (dT) primer is used in combination with a 5’ gene-specific primer to amplify 3’ cDNA ends through one, or better two, rounds of PCR. B) 5’ RACE. RNA is first decapped to free the 5’ end. In a second step, an adapter RNA sequence is added to the 5’ end. cDNA is generated by using a primer complementary to the adapter sequence. The same primer is used in combination with 3’ gene-specific primer/s to amplify the cDNA.

Figure 2. A) RACE allows amplification of several transcripts from the same gene, exemplified by 3’ RACE of Drosophila dSmad2. Combination of a nonspecific primer (Oligo dT) with a gene-specific primer (GSP) yields cDNAs of different lengths (Product A, Product B) corresponding to alternatively spliced transcripts (mRNA A, mRNA B). B) Separation of PCR products of dSmad2 3’ RACE on a 1% agarose gel. Lanes correspond to (1) 1 kb DNA ladder as marker, (2) no RT negative control (contains RNA and primers), (3) no primers negative control (contains cDNA template) (4) full reaction of 3’ RACE for dSmad2 (contains. primers and cDNA template). Amplification of dSmad2 transcripts by 3’ RACE yields three distinct products (arrows). Two cDNAs were of predicted sizes around 1400 and 1200bp (black arrows A and B, corresponding to Product A and B in (A)). In addition, a previously undescribed smaller cDNA of ~750bp was detected (grey arrow, C).
Application and Summary
RACE provides a quick, inexpensive and powerful tool to obtain cDNAs from sequences that are only partially known, or from rare transcripts that are otherwise harder to amplify. It can be used to either find the sequence of unknown transcripts from an already known gene or to clone such transcripts for further studies. Following RACE, such transcripts can be overexpressed in cell-based systems or model organisms to investigate their function in vivo.
References
- Bird, J.G., Zhang, Y., Tian, Y., Panova, N., Barvík, I., Greene, L., Liu, M., Buckley, B., Krásný, L., Lee, J.K., et al. (2016). The mechanism of RNA 5′ capping with NAD+, NADH and desphospho-CoA. Nature 535, 444–447.
- Frohman, M.A. (1994). On beyond classic RACE (rapid amplification of cDNA ends). PCR Methods Appl. 4, S40–S58.
- Frohman, M.A., Dush, M.K., and Martin, G.R. (1988). Rapid production of full-length cDNAs from rare transcripts: amplification using a single gene-specific oligonucleotide primer. Proc. Natl. Acad. Sci. U.S.a. 85, 8998–9002.
- Liu, X., and Gorovsky, M.A. (1993). Mapping the 5′ and 3′ ends of Tetrahymena thermophila mRNAs using RNA ligase mediated amplification of cDNA ends (RLM-RACE). Nucleic Acids Res. 21, 4954–4960.
- Ohara, O., Dorit, R.L., and Gilbert, W. (1989). One-sided polymerase chain reaction: the amplification of cDNA. Proc. Natl. Acad. Sci. U.S.A. 86, 5673–5677.
- Scotto Lavino, E., Du, G., and Frohman, M.A. (2007a). 3′ End cDNA amplification using classic RACE. Nat Protoc 1, 2742–2745.
- Scotto Lavino, E., Du, G., and Frohman, M.A. (2007b). 5′ end cDNA amplification using classic RACE. Nat Protoc 1, 2555–2562.
- Upadhyay, A., Moss-Taylor, L., Kim, M.-J., Ghosh, A.C., and O’Connor, M.B. (2017). TGF-β Family Signaling in Drosophila. Cold Spring Harb Perspect Biol 9, a022152.
Tags
此集合中的视频:

Now Playing
Rapid Amplification of cDNA Ends
Genetics
14.7K Views

概述了遗传分析
Genetics
40.1K Views

遗传的十字架
Genetics
59.5K Views

遗传的屏幕
Genetics
29.7K Views

遗传学和疾病的概述
Genetics
49.6K Views

单核苷酸多态性基因分型
Genetics
72.7K Views

细胞遗传学
Genetics
42.1K Views

概述了基因表达
Genetics
74.6K Views

表达谱芯片
Genetics
33.7K Views

RNA 的序列
Genetics
71.0K Views

表观遗传学的概述
Genetics
48.5K Views

DNA 甲基化分析
Genetics
26.5K Views

染色质免疫沉淀
Genetics
48.1K Views

概述了基因工程
Genetics
38.6K Views

重组和基因打靶
Genetics
28.7K Views
See More
版权所属 © 2025 MyJoVE 公司版权所有,本公司不涉及任何医疗业务和医疗服务。